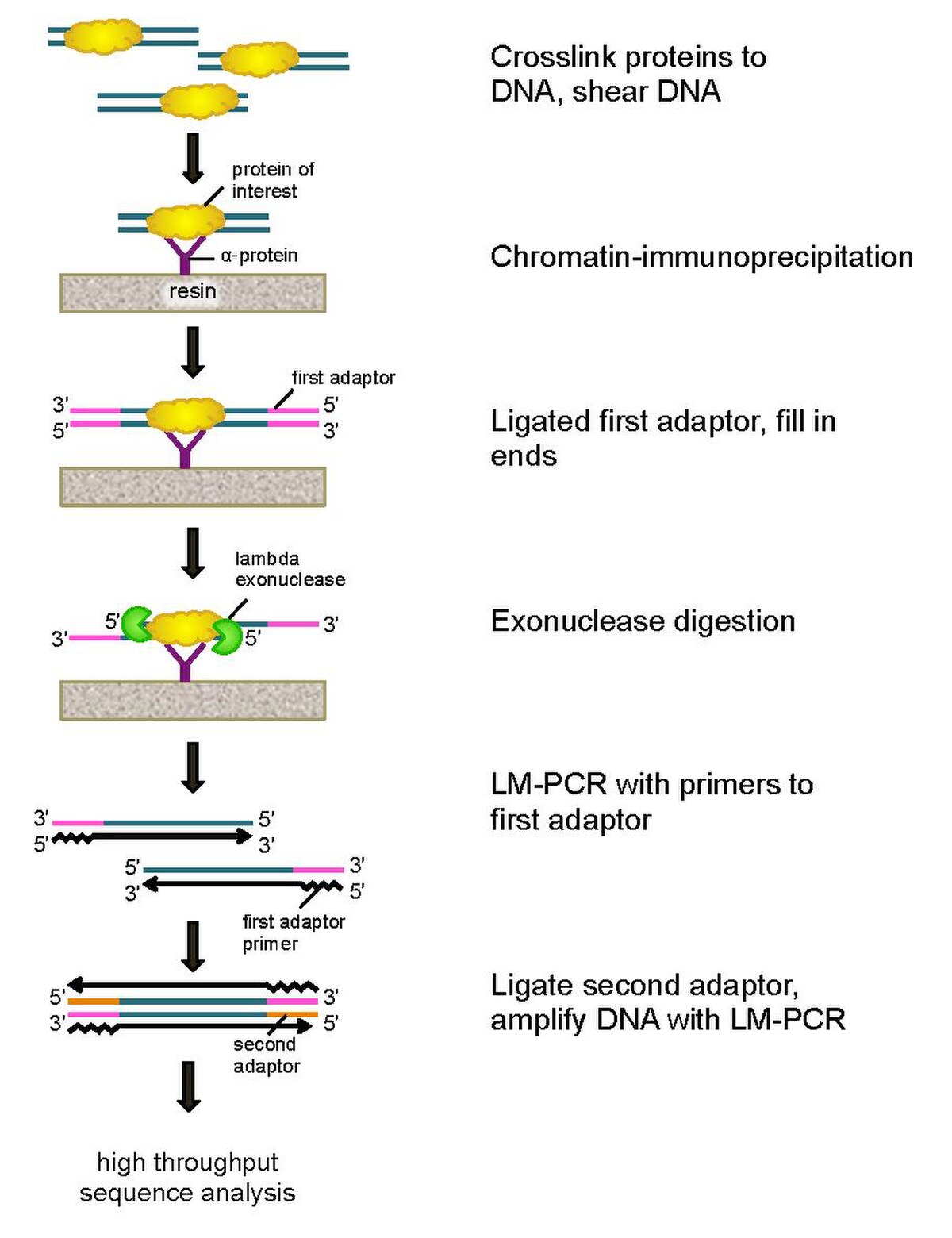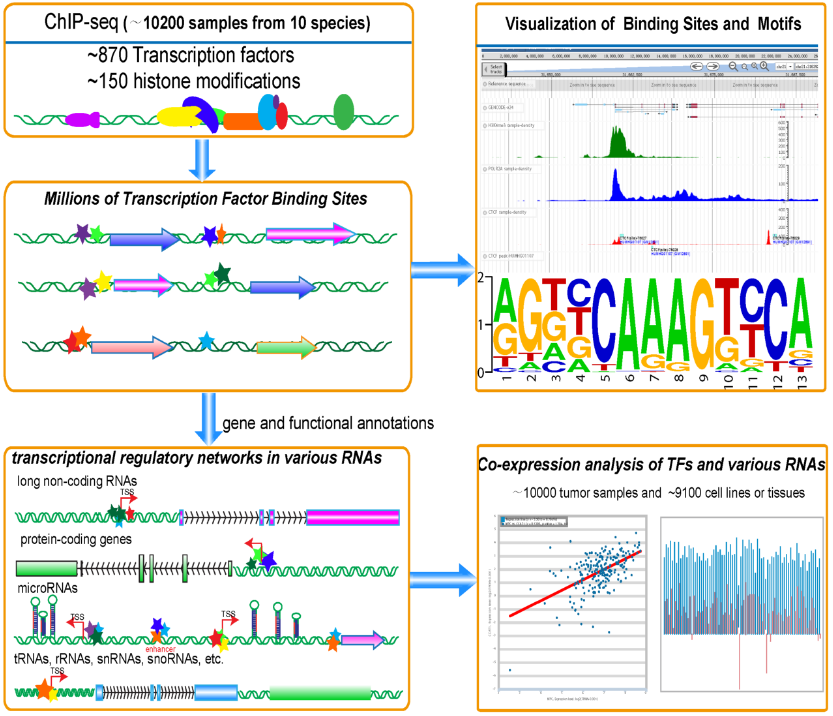
Transcription Factor Binding Sites, Motifs and Expression Profiles from ~10200 ChIP-seq and ~20000 RNA-seq samples
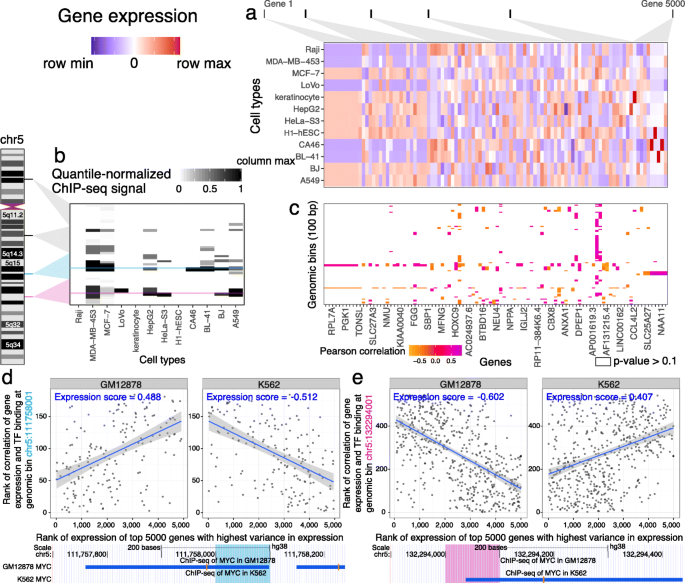
Virtual ChIP-seq: predicting transcription factor binding by learning from the transcriptome | Genome Biology | Full Text
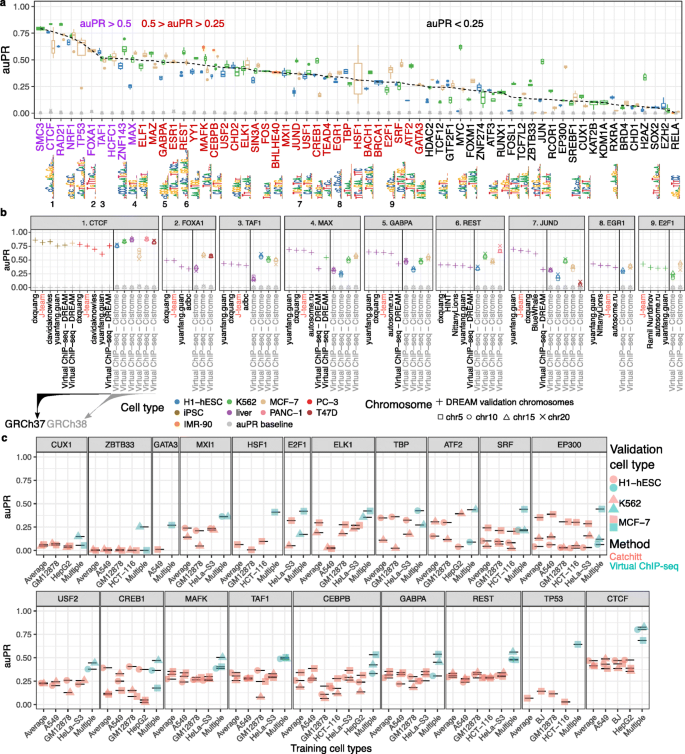

Virtual ChIP-seq: predicting transcription factor binding by learning from the transcriptome | Genome Biology | Full Text

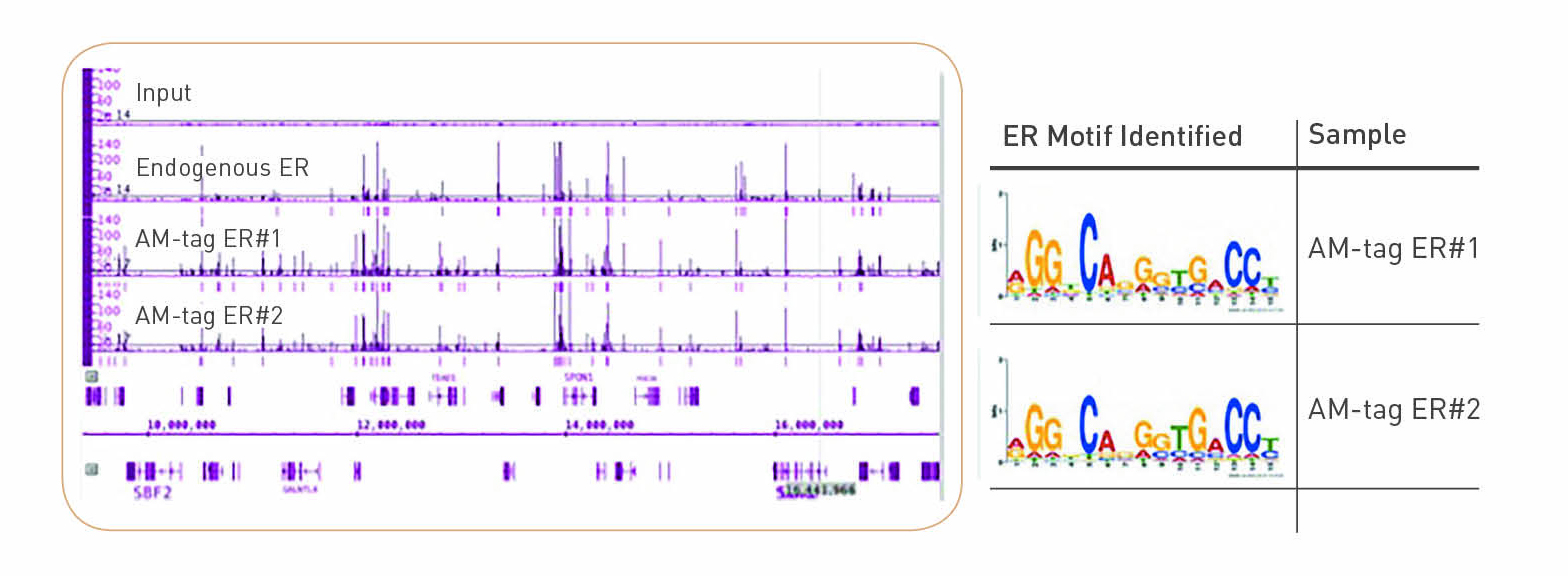
Validation of small-scale transcription factor ChIP-seq. (A) Profiles... | Download Scientific Diagram
Detection of cooperatively bound transcription factor pairs using ChIP-seq peak intensities and expectation maximization | PLOS ONE
Assessing Computational Methods for Transcription Factor Target Gene Identification Based on ChIP-seq Data | PLOS Computational Biology

10,000 cell histone mark and transcription factor ChIP-seq. (A and B)... | Download Scientific Diagram

ChIP-Seq of transcription factors predicts absolute and differential gene expression in embryonic stem cells | PNAS

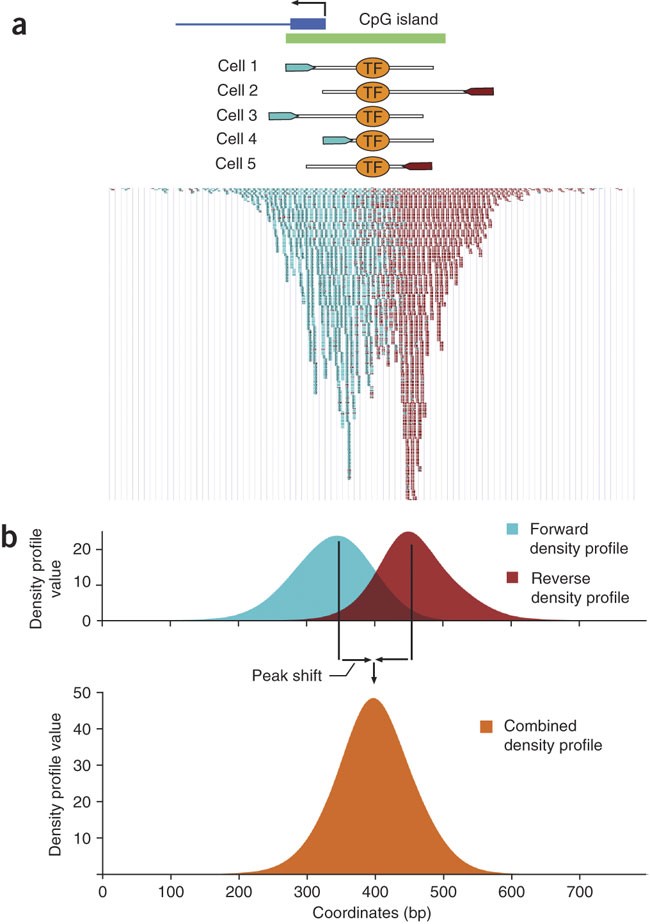
Figure 1 from Identification of transcription factor binding sites from ChIP-seq data at high resolution | Semantic Scholar