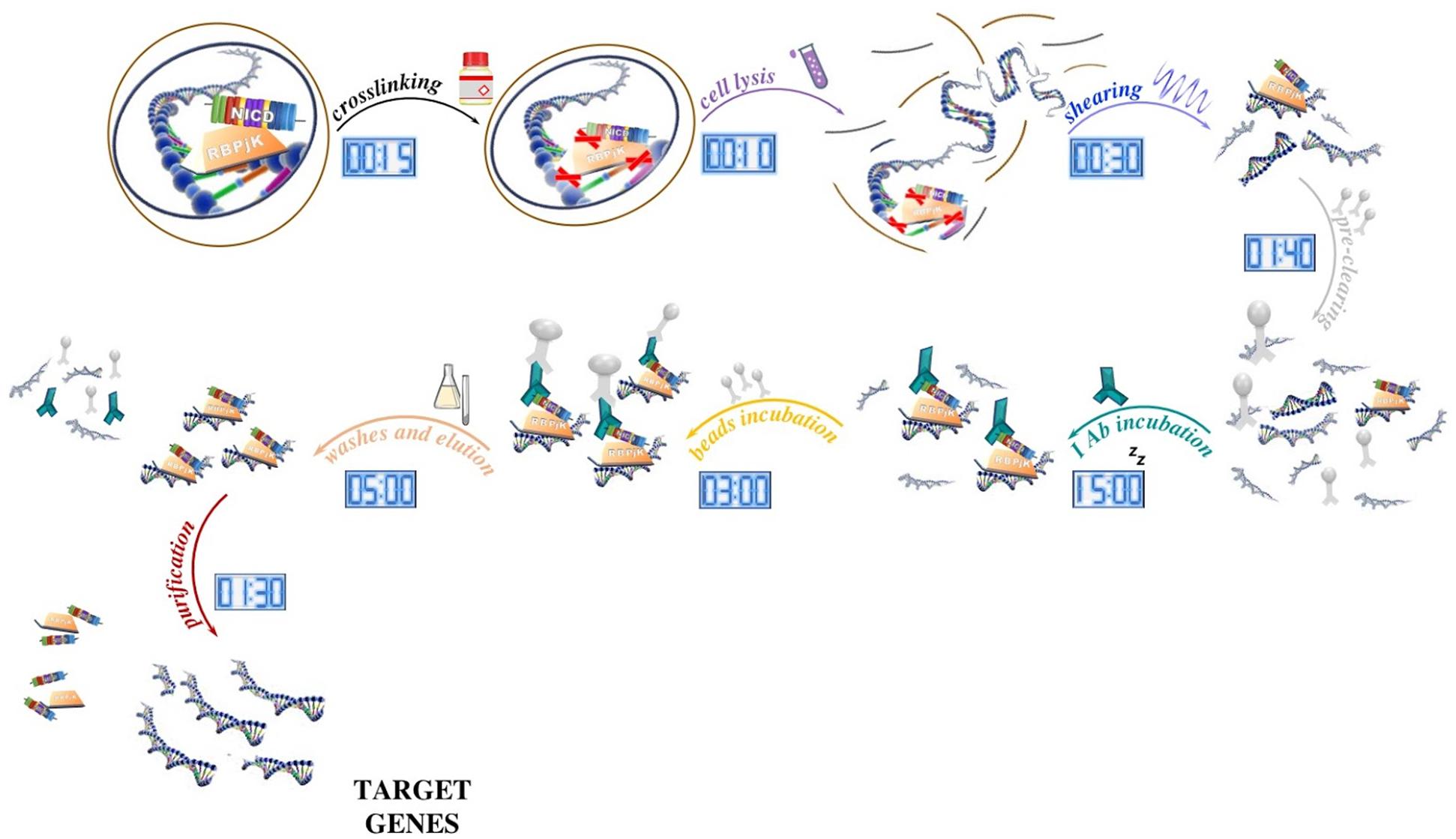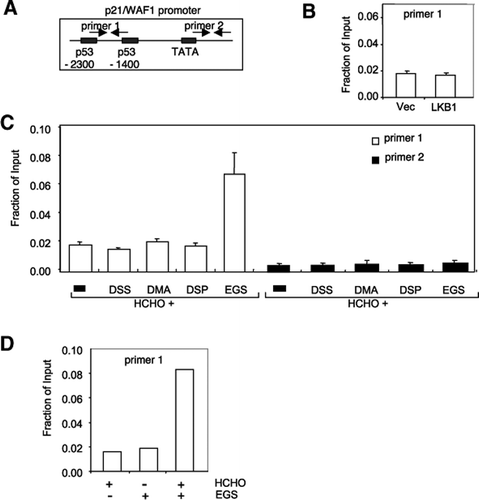
In vivo dual cross-linking for identification of indirect DNA-associated proteins by chromatin immunoprecipitation | BioTechniques
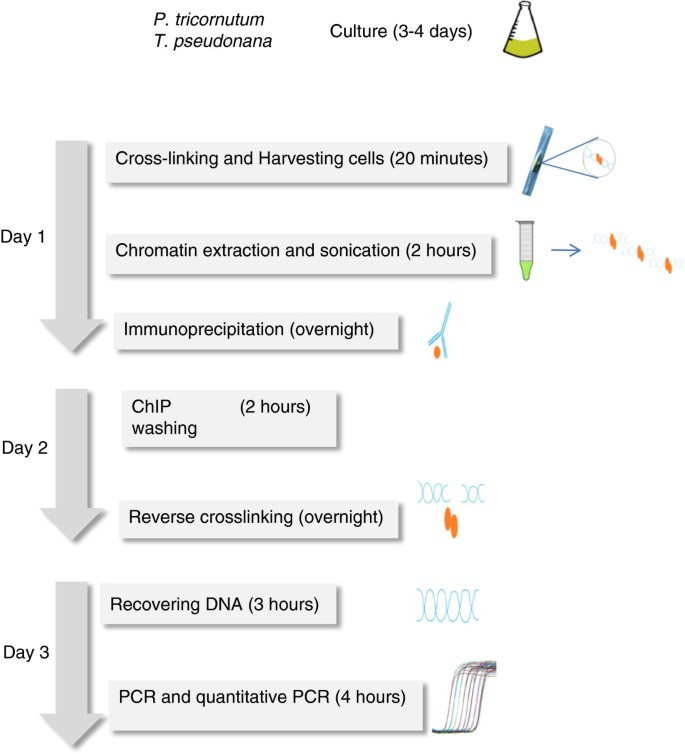
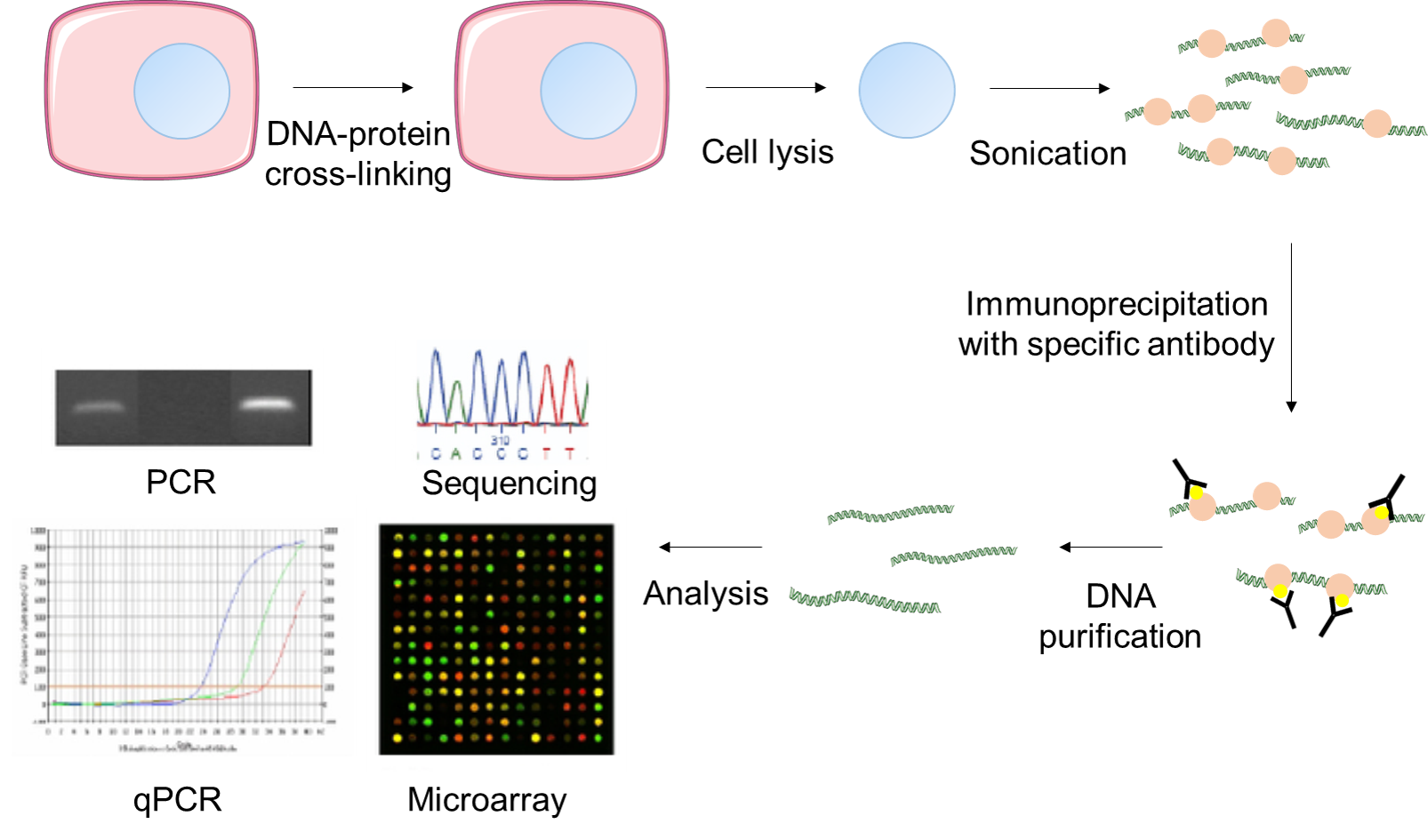
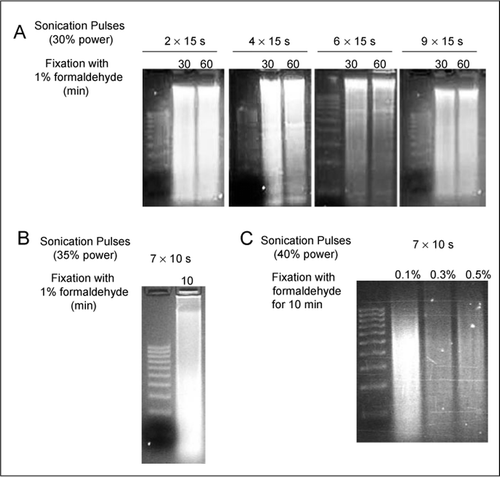
Protocol: Chromatin immunoprecipitation (ChIP) methodology to investigate histone modifications in two model diatom species | Plant Methods | Full Text

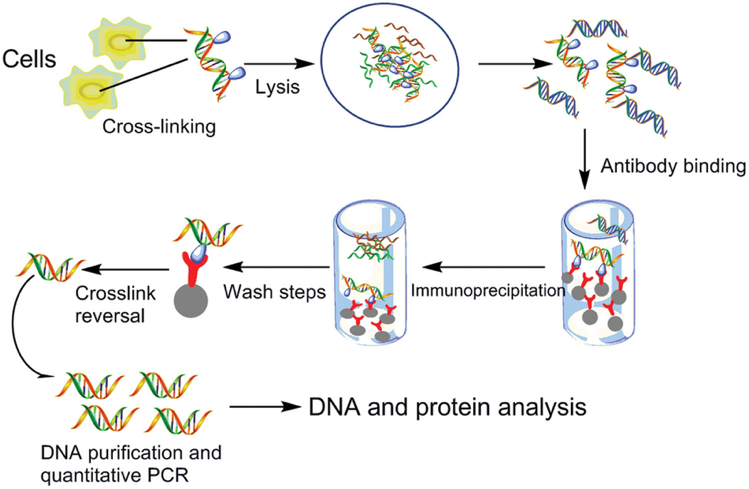
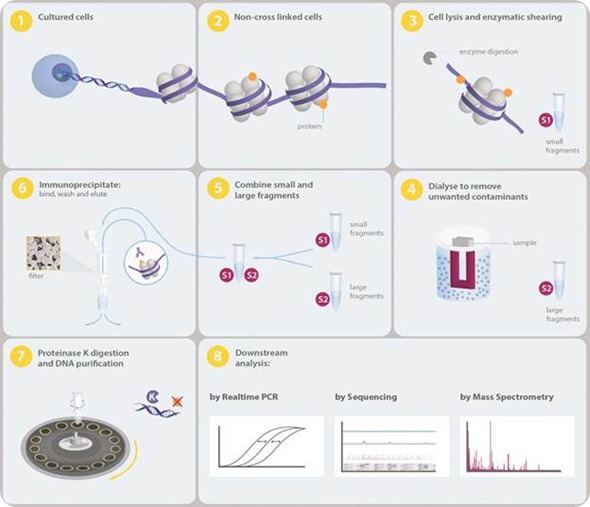
Schematic representation of the ChiP protocol. The experimental steps,... | Download Scientific Diagram

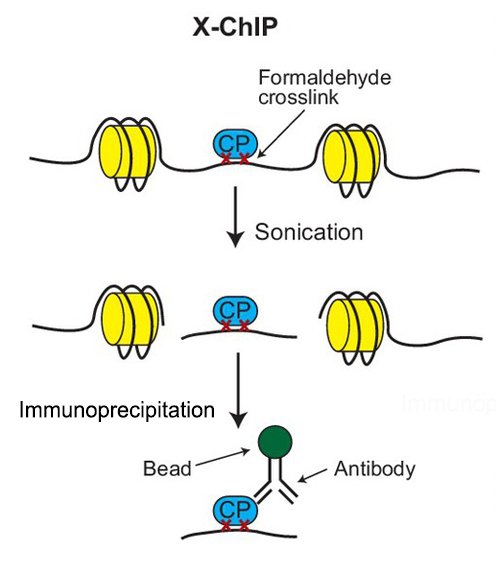
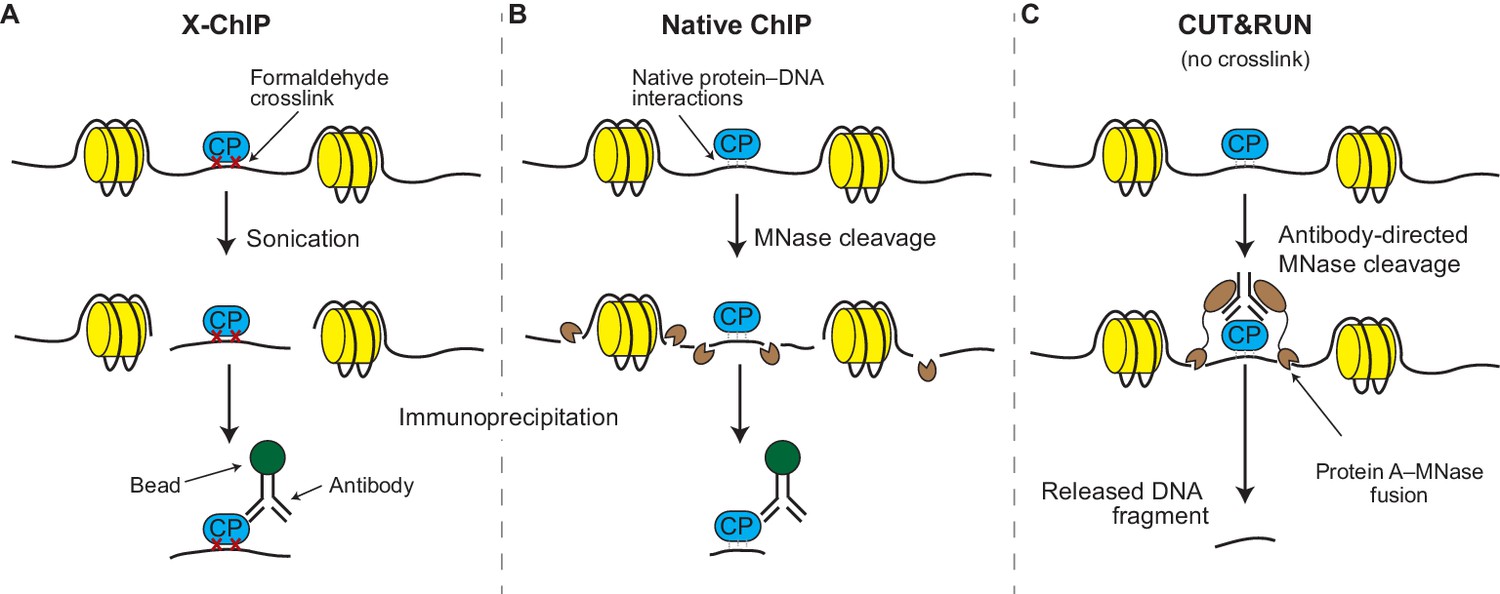
Mapping chromosomal proteins in vivo by formaldehyde-crosslinked-chromatin immunoprecipitation: Trends in Biochemical Sciences

ChIP-on-chip protocol for genome-wide analysis of transcription factor binding in Drosophila melanogaster embryos | Nature Protocols
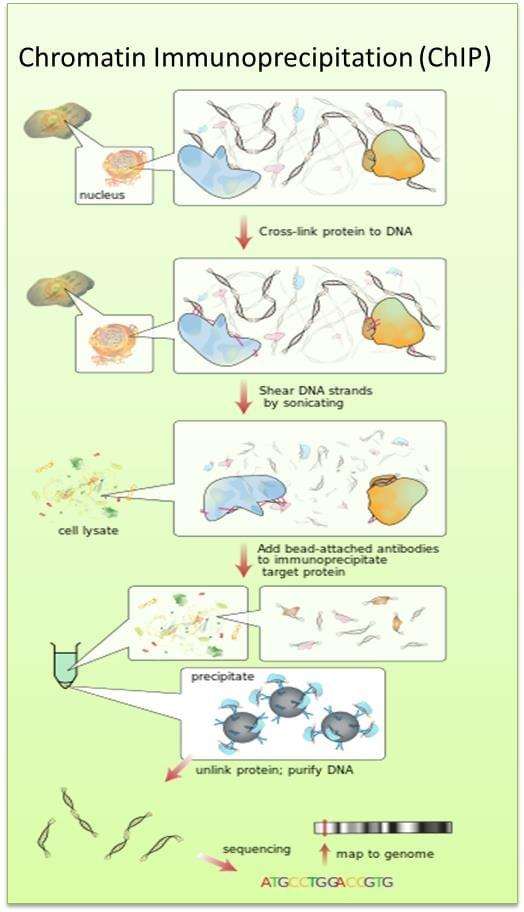
Chromatin Composition Is Changed by Poly(ADP-ribosyl)ation during Chromatin Immunoprecipitation | PLOS ONE

Scientific Protocols - Combined 3C-ChIP-Cloning (6C) Assay: A Tool to Unravel Protein-Mediated Genome Architecture

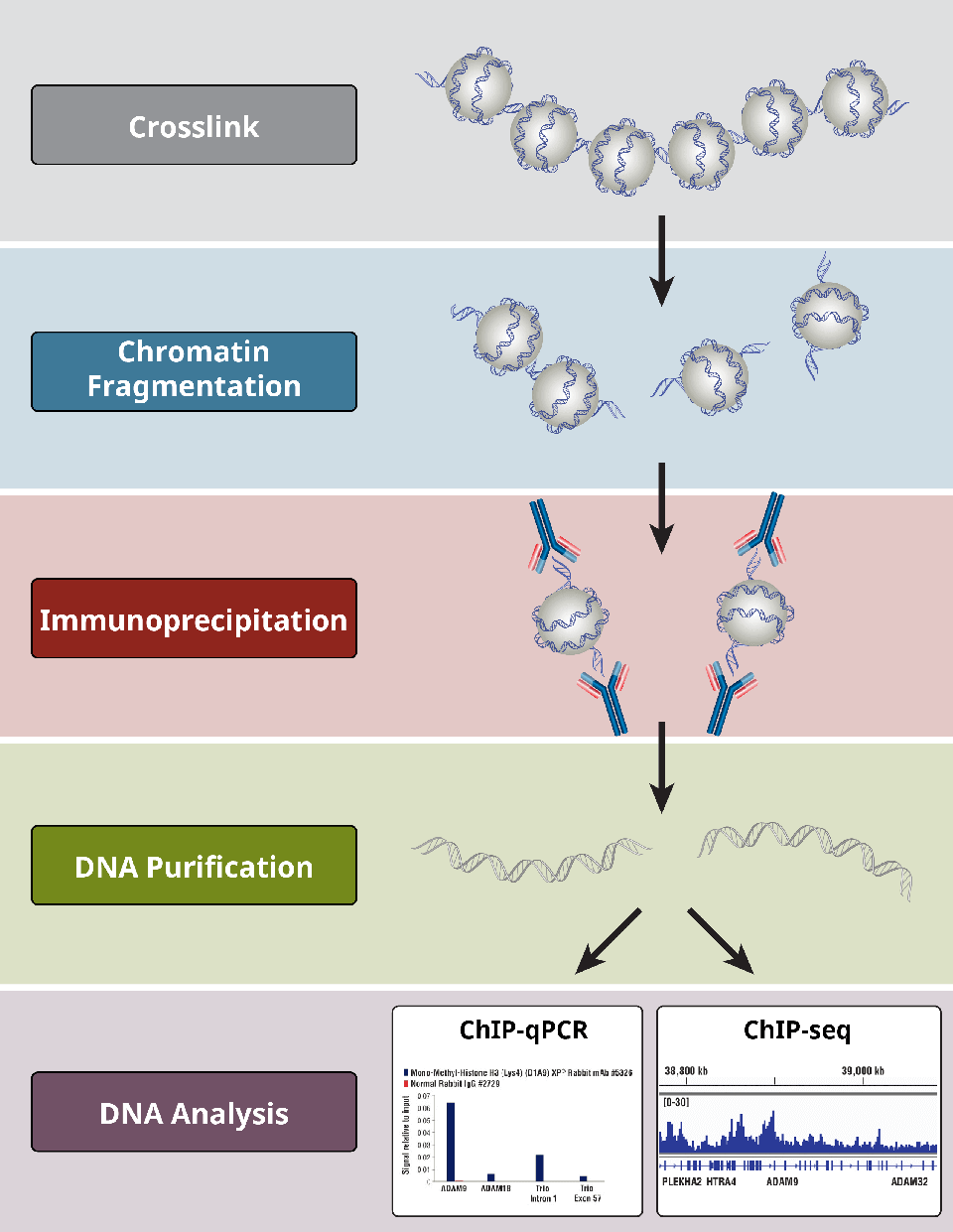
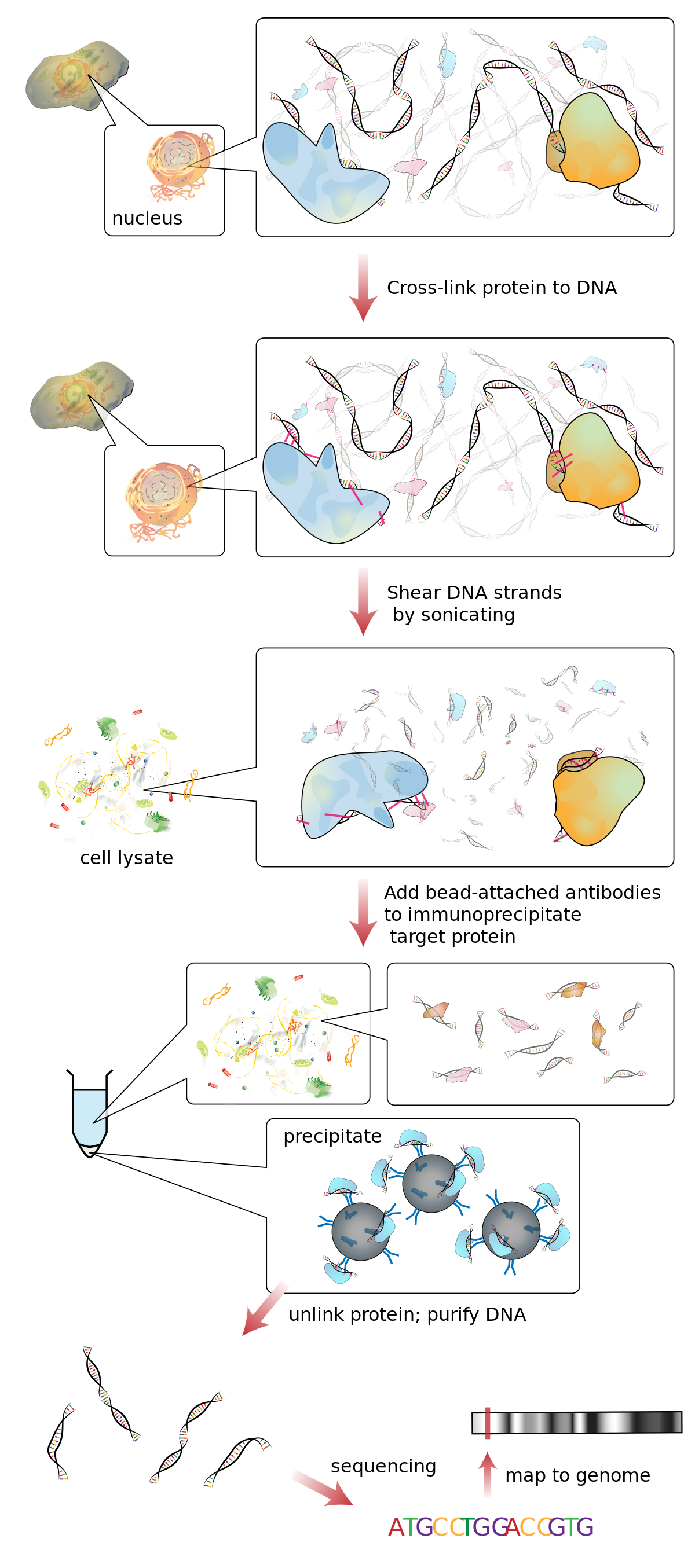
Outline of the ChIP procedure. After cross-linking with formaldehyde,... | Download Scientific Diagram